A key feature of therapeutic IVT mRNAs is that they contain modified ribonucleotides, which have been shown to decrease innate immunogenicity and can additionally increase mRNA stability, both of which are favourable characteristics for mRNA therapies1,2,3,4,5. For example, clinically approved SARS-CoV-2 mRNA vaccines incorporate N1-methylpseudouridine (1-methylΨ), which has been shown to decrease IVT mRNA innate immunogenicity3,4,5. Some modified ribonucleotides, such as 5-methylcytidine (5-methylC), are naturally occurring post-transcriptional mRNA modifications in eukaryotes, whereas others are not, such as 1-methylΨ (refs. 6,7,8,9,10).
We investigated how 5-methoxyuridine (5-methoxyU), 5-methylC and 1-methylΨ affect translation of IVT mRNA. 5-methoxyU, 5-methylC and 1-methylΨ have been utilized in IVT mRNAs to attempt to increase recombinant protein synthesis in vitro, and for preclinical proof of concept for IVT mRNA-based therapies11,12. As mentioned, 1-methylΨ is a ribonucleotide incorporated in licensed IVT mRNA-based SARS-CoV-2 vaccines, but also mRNA-based human vaccines and therapies in development2,13,14.
Despite their widespread use, surprisingly little is known about how ribonucleotide modification affects protein synthesis, particularly for translation of therapeutic IVT mRNAs. We were interested in how modified ribonucleotides affect the fidelity of mRNA translation for several reasons. Certain ribonucleotide modifications can recode mRNA sequences (for example, inosine15). 5-methylC has previously been shown to increase misreading during mRNA translation in prokaryotes, but its effect on eukaryotic mRNA translation fidelity has not been explored16. The effect of 5-methoxyU on translation fidelity has not been investigated. Pseudouridine (Ψ) is known to increase misreading of mRNA stop codons in eukaryotes, and can affect misreading during prokaryotic mRNA translation16,17,18. 1-methylΨ does not seem to affect codon misreading, but has been shown to affect protein synthesis rates and ribosome density on mRNAs, suggesting a direct effect on mRNA translation19,20.
At present, it is unclear which modified ribonucleotides affect mRNA translation fidelity and existing studies are mostly limited to understanding misreading frequencies only at a given codon. Misreading of mRNA codons is also only one type of post-transcriptional mechanism that can alter a polypeptide sequence. So far, no study has investigated the fundamental question of whether modified ribonucleotides can affect the maintenance of the correct reading frame during translation of a synthetic transcript. Understanding these processes is critical to increase our knowledge of protein synthesis from modified mRNAs in general, but is also imperative for the robust design and evaluation of new mRNA-based therapeutics that make use of modified ribonucleotides within widely differing RNA sequences or therapeutic contexts.
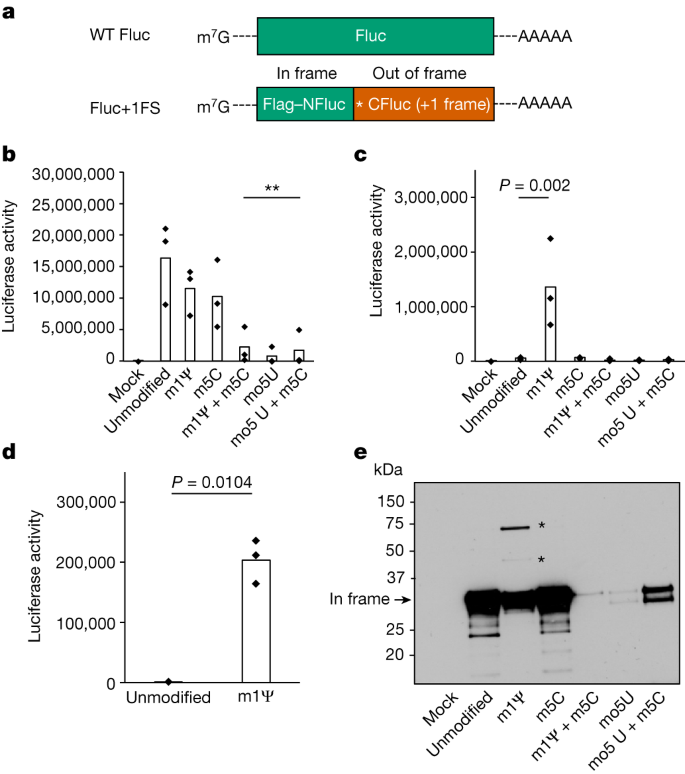
To investigate how ribonucleotide modification affects reading frame maintenance during translation of mRNA, we designed and synthesized IVT mRNAs (Fluc+1FS) that report on out-of-frame protein synthesis (Fig. 1a). Fluc+1FS mRNAs encode an amino-terminal segment of firefly luciferase (NFluc) and a complementary carboxy-terminal segment of Fluc (CFluc), directly downstream. CFluc is encoded in the +1 reading frame. Fluc+1FS mRNAs are designed to produce catalytically inactive (truncated) NFluc when translated normally. However, if ribosomes move out of frame during translation, elongated polypeptides containing residues from both in-frame NFluc and out-of-frame CFluc can be produced, which can increase catalytic activity.
a, Structures of IVT mRNA transcripts used to probe protein synthesis fidelity. WT Fluc contains only (in-frame) Fluc coding sequence. For Fluc+1FS, the green segment represents in-frame N-terminal Fluc coding sequence (NFluc), and the orange segment represents +1 frameshifted C-terminal Fluc coding sequence (CFluc). Asterisk represents a premature stop codon. b, Luciferase activity produced by translation of WT Fluc mRNAs, either unmodified control (canonical nucleotides), or containing 1-methylΨ (m1Ψ), 5-methylC (m5C), 5-methoxyU (mo5U) or the combinations indicated. **P < 0.01 (1-methylΨ + 5-methylC, P = 0.0051; 5-methoxyU, P = 0.0023; 5-methoxyU + 5-methylC, P = 0.0042; one-way analysis of variance (ANOVA) with Dunnett’s test). c, Luciferase activity produced by translation of modified Fluc+1FS mRNAs and unmodified control. 1-methylΨ, P = 0.002 (one-way ANOVA with Dunnett’s test). d, Luciferase activity in lysates produced by transfection of HeLa cells with unmodified or 1-methylΨ Fluc+1FS mRNA for 8 h. P = 0.0104 (Welch’s one-tailed t-test). e, Western blot analysis (anti-Flag epitope) of polypeptides produced by translation of mRNAs in c. All data are obtained from n = 3 replicated experiments. e shows a single blot from n = 3 replicated experiments. Asterisks represent bands at higher molecular weight. For gel source data, see Supplementary Fig. 2.
We synthesized unmodified Fluc+1FS mRNAs, which contain canonical ribonucleotides, and translated them in vitro. We confirmed that Fluc+1FS mRNAs produce catalytically inactive NFluc (Extended Data Fig. 1). By comparison, unmodified wild-type (WT) Fluc mRNA, containing the complete in-frame Fluc coding sequence, produced the expected active protein (Extended Data Fig. 1). Then we synthesized and translated each mRNA containing 5-methoxyU, 5-methylC, 1-methylΨ, 5-methoxyU + 5-methylC or 1-methylΨ + 5-methylC. Translation of WT Fluc mRNA was not significantly affected by either 1-methylΨ or 5-methylC modifications alone, but was decreased by incorporating both ribonucleotides into a single transcript (Fig. 1b). 5-methoxyU incorporation alone, or combined with 5-methylC, significantly decreased translation of WT Fluc mRNA (Fig. 1b). Incorporation of 1-methylΨ in Fluc+1FS mRNA significantly increased ribosomal +1 frameshifting to about 8% of the corresponding in-frame protein, which was not observed for other ribonucleotides (Fig. 1c). HeLa cells transfected with 1-methylΨ Fluc+1FS mRNA recapitulated the results from in vitro translation (Fig. 1d). On the basis of these observations, we concluded that IVT mRNA containing 1-methylΨ or 5-methylC exhibits similar translation efficiency to unmodified mRNA, but 1-methylΨ significantly increases ribosomal +1 frameshifting during mRNA translation.
We observed a large increase in ribosomal +1 frameshifting during translation of 1-methylΨ mRNA and reasoned that gaining better understanding of the translation products would complement the reporter assay data and help to explain how +1 frameshifted products originate. To address these aspects, we probed the polypeptides produced during IVT mRNA translation by western blotting. Translation of unmodified Fluc+1FS mRNA produced the expected in-frame truncated product, which was also true for 5-methylC mRNA (Fig. 1e). Translation of 1-methylΨ mRNA produced the expected in-frame product, but also produced two additional bands at higher molecular weight (Fig. 1e). We reasoned that these products were +1 frameshifted polypeptides. We also confirmed that 1-methylΨ + 5-methylC-, 5-methoxyU- and 5-methoxyU + 5-methylC mRNAs were comparatively poor mRNA templates for protein synthesis (Fig. 1e).
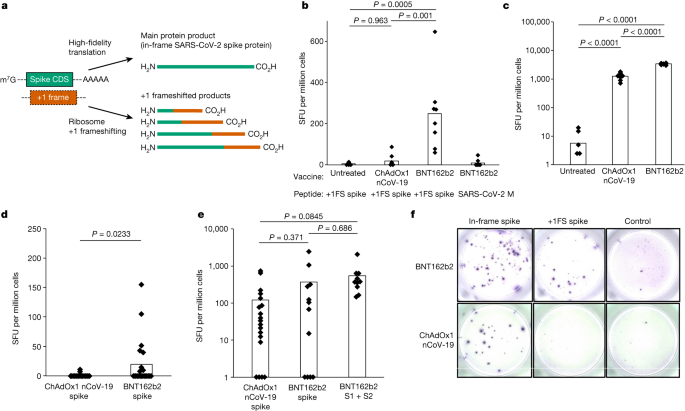
1-methylΨ is also used in clinically approved SARS-CoV-2 mRNA vaccines3,4. As 1-methylΨ increased +1 ribosome frameshifting during translation in vitro, we investigated whether this occurs in vivo for BNT162b2, a SARS-CoV-2 mRNA vaccine containing 1-methylΨ. We reasoned that +1 ribosomal frameshifting during recombinant antigen mRNA translation could lead to presentation of +1 frameshifted products to T cells, and elicit off-target cellular immune responses (Fig. 2a). Antigen presentation from mistranslation of endogenous tumour mRNA has been shown to occur in vivo (for example, ref. 21). To address this possibility, we vaccinated mice with BNT162b2 and quantified their T cell response to in-frame SARS-CoV-2 spike protein and +1 frameshifted products predicted to occur by translation of the mRNA +1 frame, as well as an unrelated control antigen (SARS-CoV-2 M protein), by interferon-γ (IFNγ) ELISpot assay. Junction peptides consisting of in-frame N-terminal residues and C-terminal +1 frameshifted residues were not included. We found that responses to +1 frameshifted spike peptides were significantly increased in vaccinated mice compared to untreated mice or those vaccinated with ChAdOx nCoV-19, which does not produce antigen from translation of N1-methylpseudouridylated mRNA22 (Fig. 2b). Both BNT162b2 and ChAdOx1 nCoV-19 vaccination produced ELISpot responses to in-frame SARS-CoV-2 spike (Fig. 2c). These data suggest that +1 frameshifted products encoded in BNT162b2 spike mRNA are T cell antigens for inbred mice, to which off-target immunity can be detected following vaccination.
a, Depiction of spike and +1 frameshifted (+1FS) products produced by 1-methylΨ-modified spike mRNA translation. CDS, coding sequence. b, Splenocyte IFNγ ELISpot responses from untreated, ChAdOx1 nCoV-19-vaccinated or BNT162b2-vaccinated mice stimulated with +1FS spike peptides. IFNγ ELISpot response from BNT162b2-vaccinated mice stimulated with SARS-CoV-2 M peptides (unrelated control antigen) is included for additional comparison. SFU, spot-forming units. Each group n = 8. Untreated versus ChAdOx1 nCoV-19, P = 0.963; untreated versus BNT162b2, P = 0.0005; ChAdOx1 nCoV-19 versus BNT162b2, P = 0.001. c, Splenocyte IFNγ ELISpot responses from mice in b stimulated with spike peptides. Untreated versus ChAdOx1 nCoV-19, P = 2.05 × 10−9; untreated versus BNT162b2, P = 4.5 × 10−14; ChAdOx1 nCoV-19 versus BNT162b2, P = 1.88 × 10−13. d, Peripheral blood mononuclear cells (PBMC) IFNγ ELISpot responses from donors vaccinated with ChAdOx1 nCoV-19 (n = 20) or BNT162b2 (n = 21) stimulated with +1FS spike peptides. P = 0.0233 (Welch’s one-tailed t-test). e, PBMC IFNγ ELISpot responses from donors in c stimulated with in-frame spike peptides: total spike pool or spike S1 + S2 subpools. ChAdOx1 nCoV-19 spike versus BNT162b2 spike, P = 0.371; ChAdOx1 nCoV-19 spike versus BNT162b2 S1 + S2, P = 0.0845; BNT162b2 spike versus BNT162b2 S1 + S2, P = 0.686. f, Representative images of PBMC IFNγ ELISpot response wells for two individuals vaccinated with either BNT162b2 responder (top) or ChAdOx1 nCoV-19 (bottom). Left to right: in-frame spike response (spike peptides); +1FS spike response (+1FS spike peptides); no peptide control. P values in b,c,e were determined by one-way ANOVA and Tukey’s test.
We then compared IFNγ ELISpot responses to predicted +1 frameshifted SARS-CoV-2 spike protein products in 21 individuals vaccinated with BNT162b2 and compared these responses to those of 20 individuals vaccinated with ChAdOx1 nCoV-19, none of whom reported undue effects as a result of vaccination. We detected a significantly higher IFNγ response to +1 frameshifted antigen in the BNT162b2 vaccine group, compared to ChAdOx1 nCoV-19 (Fig. 2d). There was no association between T cell responses to +1 frameshifted antigen and age, sex or HLA subtype (Supplementary Table 1 and Extended Data Figs. 2 and 3). Both ChAdOx1 nCoV-19 and BNT162b2 vaccination produced ELISpot responses to in-frame SARS-CoV-2 spike, but responses to +1 frameshifted products were observed only in individuals vaccinated with BNT162b2 (Fig. 2e,f). During SARS-CoV-2 viral replication, a programmed −1 ribosomal frameshift occurs naturally during translation of open reading frame (ORF) 1a and ORF1b (ref. 23). It is not feasible that these data are a consequence of natural SARS-CoV-2 infection for the following, non-exhaustive, reasons. First, no frameshifting activity is known to occur during SARS-CoV-2 spike subgenomic mRNA translation (which would be a major discovery in its own right). Second, −1 frameshifting (and not +1 frameshifting) is restricted to a single programmed site in ORF1a and ORF1b (ref. 23). Third, +1 frameshifted peptides are predicted from the BNT162b2 mRNA sequence, and not the S gene sequence from wild virus (Extended Data Fig. 4). Instead, these data suggest that vaccination with 1-methylΨ mRNA can elicit cellular immunity to peptide antigens produced by +1 ribosomal frameshifting in both major histocompatibility complex (MHC)-diverse people and MHC-uniform mice.
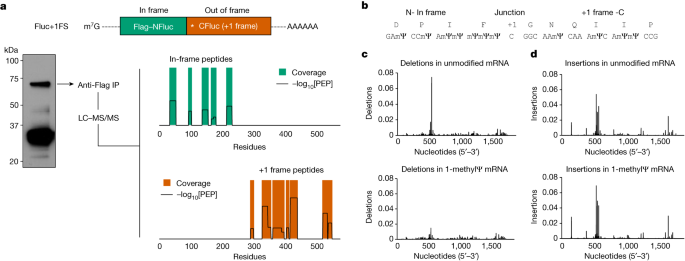
To provide further mechanistic insight into +1 ribosome frameshifting during translation of 1-methylΨ mRNA, and identify potential frameshift sites or sequences, we translated 1-methylΨ Fluc+1FS mRNA, purified the major putative +1 frameshifted polypeptide and carried out liquid chromatography tandem mass spectrometry (LC–MS/MS) of tryptic digests. From this single polypeptide, we identified six in-frame peptides and nine peptides derived from the mRNA +1 frame (Fig. 3a and Extended Data Table 1). All in-frame peptides were mapped to the N-terminal region, whereas +1 frameshifted peptides were mapped downstream (Fig. 3a). We then repeated this analysis using a different protease and identified a junction peptide spanning the main frame and the +1 frame (Fig. 3b). These data demonstrated that the elongated polypeptide was indeed a chimeric polypeptide consisting of in-frame N-terminal residues and +1 frameshifted C-terminal residues. As expected, shorter frameshifted products were also produced from translation of 1-methylΨ mRNA encoding full-length Fluc (Extended Data Fig. 5).
a, Tryptic peptide coverage plot of the purified high molecular weight polypeptide produced by translation of 1-methylΨ Fluc+1FS mRNA, showing in-frame residues (top) and +1 frameshifted residues (bottom). −log10[PEP] is the mass spectrum percolator score (only high-quality peptides are shown). IP, immunoprecipitate. The structure of Fluc+1FS mRNA from Fig. 1 is re-displayed and a western blot of the translation reaction before immunoprecipitation is displayed. For gel source data, see Supplementary Fig. 3. b, Junction peptide derived from +1 ribosomal frameshifting and the originating mRNA sequence. c, Nucleotide deletions in unmodified (top) and 1-methylΨ (bottom) Fluc+1FS mRNA, quantified by n = 3 RNA-sequencing analyses. d, Nucleotide insertions in unmodified (top) and 1-methylΨ (bottom) Fluc+1FS mRNA.
Apparent errors in protein synthesis, including frameshifting, can be consequences of DNA mutation or transcriptional errors24. Hence, faithful translation of an incorrect mRNA sequence can produce incorrect proteins. In vitro transcripts are presumed to be exact RNA copies of template DNA, the accuracy of which may be estimated by the fidelity of the used RNA polymerase. However, the substitution of canonical substrate ribonucleoside triphosphates for modified nucleotides may increase transcriptional errors. To address this possibility, we carried out high-throughput RNA sequencing of unmodified and 1-methylΨ Fluc+1FS mRNA and quantified nucleotide insertions and deletions in each population of IVT mRNA. Nucleotide deletion profiles for each mRNA were very similar (Fig. 3c), as were nucleotide insertions (Fig. 3d), suggesting few site-specific differences. The overall frequency of insertions and deletions was low, and did not differ significantly between unmodified and 1-methylΨ mRNA (Extended Data Table 2), which is supported by recent observations25. From these findings, we concluded that frameshifted products of 1-methylΨ mRNA translation were not due to transcriptional errors, but were due to bona fide ribosomal +1 frameshifting—a post-transcriptional mechanism.
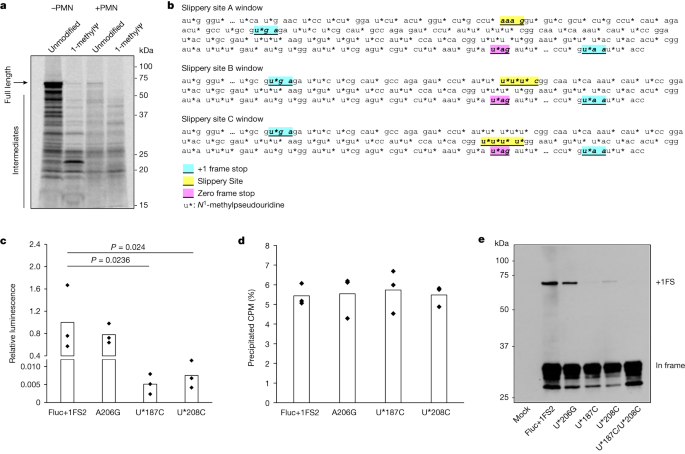
Ribosome frameshifting is a well-documented phenomenon that occurs during translation of many naturally occurring mRNAs24. As ribosome stalling is implicated in several instances of +1 frameshifting, we queried how the presence of 1-methylΨ in IVT mRNA affects translation elongation26,27,28. To do this, we assayed protein synthesis during translation of unmodified or 1-methylΨ WT Fluc mRNA using co-translational [35S]methionine labelling29. Translation elongation of 1-methylΨ mRNA was slower than for unmodified mRNA (Fig. 4a), which is supported by previous observations20. All reactions were run for 30 min and there was less full-length protein produced from the translation of 1-methylΨ-containing mRNAs, suggesting a slower elongation rate compared to that of unmodified mRNA, with a greater proportion of premature polypeptide products. These data suggested that elongating ribosomes stall during translation of mRNA containing 1-methylΨ.
a, SDS–polyacrylamide gel electrophoresis autoradiograph of [35S]methionine-labelled polypeptides produced by translation of unmodified or 1-methylΨ Fluc mRNA for 30 min, including or omitting 100 μM paromomycin (+PMN and −PMN, respectively). b, Diagram showing putative mRNA slippery sequences and stop-codon-flanked windows. c, Activity of +1 frameshifted products after translation of 1-methylΨ mutant mRNAs, or 1-methylΨ Fluc+1FS2 control mRNA, for 2 h. Fluc+1FS2 versus U*187C, P = 0.024; Fluc+1FS2 versus U*208C, P = 0.0236 (one-way ANOVA with Dunnett’s test). d, Total mRNA translation over 2 h for each of Fluc+1FS2 mRNA or mutant mRNAs, quantified by [35S]methionine incorporation. CPM, counts per minute. e, Western blot analysis (anti-Flag epitope) of polypeptides produced by translation of mRNAs in c, and U*187C/U*208C double-mutant 1-methylΨ mRNA. Data are from n = 3 replicated experiments. a and e show representative images from n = 3 replicated experiments. For gel source data, see Supplementary Figs. 4 and 5.
It was unclear whether 1-methylΨ affected mRNA decoding rates, or another process, during elongation. We reasoned that slower decoding of 1-methylΨ codons during translation elongation could lead to ribosome stalling, similar to previous observations for ‘hungry’ codons at sites of +1 frameshifting during translation of naturally occurring mRNA21,28. We probed the molecular mechanism of ribosome stalling during 1-methylΨ mRNA translation using the aminoglycoside paromomycin. In brief, during mRNA decoding, cognate aminoacyl-tRNA anticodon–codon interaction causes local conformational changes in 18S rRNA (in eukaryotes), after which a new peptide bond is formed, ribosome subunit rotation occurs, and subsequent ribosome conformational changes, elongation factor 2 binding and translocation to the next codon completes the elongation cycle30. Paromomycin binds to helix 44 of 18S rRNA in elongating ribosomes and alters its conformation in the decoding centre, which inhibits translation but also permits the productive binding of near- and non-cognate aminoacyl-tRNAs to the 80S ribosome A-site31. In doing so, paromomycin increases the misincorporation of amino acids into elongating polypeptides32. We reasoned that if slow decoding during 1-methylΨ mRNA translation was due to altered aminoacyl-tRNA binding kinetics, this process could be decreased by paromomycin. This is because paromomycin-bound ribosomes could incorporate additional near- or non-cognate aminoacyl-tRNAs and effectively increase the substrate aminoacyl-tRNA pool at ribosome stall sites. Translation of 1-methylΨ mRNA was slower than that of unmodified mRNA and the proportion of premature polypeptide products was greater (Fig. 4a). However, during 1-methylΨ mRNA translation, polypeptide elongation was improved by the addition of paromomycin, whereas paromomycin was inhibitory only to unmodified mRNA translation (Fig. 4a). Together, these data show that slow translation of 1-methylΨ mRNA is probably due to ribosome stalling, which is caused by altered aminoacyl-tRNA binding, and which can be rescued by increasing the incorporation of near- or non-cognate amino acids into elongating polypeptides.
Although there is no evidence that frameshifted products in humans generated from BNT162b2 vaccination are associated with adverse outcomes, for future use of mRNA technology it is important that mRNA sequence design is modified to reduce ribosome frameshifting events, as this may limit its future use for applications that require higher doses or more frequent dosing, such as the in vivo production of hormones. It is important to continue investigating therapeutic mRNA mistranslation and immunogenicity, as the evolution of antibody and cytolytic T cell responses against +1 frameshifted spike variants and peptides has not been systematically evaluated in humans and ELISpot responses obtained from pooled peptides may also underestimate T cell responses. The main in-frame mRNA-encoded product is unlikely to elicit an adaptive immune response, but presentation of +1 frameshifted products could activate T cells that target host cells. We reasoned that if we were able to identify +1 ribosome frameshift sites or sequences it would be possible to alter the mRNA sequence to reduce such effects. As proof of principle, we used our reporter IVT mRNA system. LC–MS/MS analysis showed that translation of 1-methylΨ mRNA leads to synthesis of +1 frameshifted products within the area of coding sequence between detected in-frame residues and downstream +1 frameshifted residues (Fig. 3a). We searched the RNA sequence corresponding to this region in the junction peptide coding sequence (Fig. 3b) and determinants of ribosome frameshifting from published mechanisms, from which we identified three potential ribosome slippery sequences (Fig. 4c), with all three sequences having the potential to be decoded by the same aminoacyl-tRNA at an in-frame codon or in the immediate +1 frame codon. Notably, six slippery sites identical to Fluc+1FS slippery sites B and C were also distributed in the BNT162b2 spike mRNA coding sequence. These sites have been annotated in the Fluc+1FS coding sequence (Fig. 4b) and the BNT163b2 spike mRNA coding sequence (Extended Data Fig. 6). We reasoned that these sequences could therefore function as sites for +1 ribosomal frameshifting. We synonymously mutated each site in 1-methylΨ Fluc+1FS mRNA such that the in-frame amino acid was unchanged, but the immediate +1 frame codon was mutated to a non-cognate amino acid, hence destroying the ribosome slippery sequence, and translated the mRNAs to evaluate the contribution of each site to +1 ribosomal frameshifting (Fig. 4c). A +1 frame stop codon was present downstream of slippery site A, and it was unlikely that frameshifting at this site contributed to increased luciferase activity. As expected, luciferase activity produced by translation of site A mutant A206G mRNA was the same as control levels (Fig. 4c). However, both slippery site B mutant U*187C mRNA and slippery site C mutant U*208C mRNA strongly decreased +1 ribosome frameshifting (Fig. 4c). Notably, translation efficiency of each mRNA was equal, which suggests that no mutation adversely affected mRNA translation overall, but solely +1 ribosomal frameshifting activity (Fig. 4d). Translation of a U*187C/U*208C double-mutant 1-methylΨ Fluc+1FS mRNA produced no detectable +1 ribosome frameshifting (Fig. 4e). The transframe protein product predicted by +1 frameshifting at slippery site C contains an alteration of 19 amino acid residues (compared to WT Fluc), whereas +1 frameshifting at slippery site B produces a transframe product that is effectively 100% homologous to WT Fluc. In addition, given that mutation of either slippery site B or C (U*187C or U*208C) significantly decreased luciferase activity, but that relatively more frameshifted product was produced by translation of U*208C mRNA (Fig. 4e), we reasoned that the transframe product produced by frameshifting at slippery site C had lower specific luciferase activity, and that frameshifting at slippery site B contributed to most of the detected luciferase activity as a consequence of +1 ribosome frameshifting. Taken together, these data suggest that N1-methylpseudouridylation at defined mRNA sequences triggers ribosome +1 frameshifting; however, with appropriate mRNA sequence design, it is possible to ameliorate this issue.
Source link